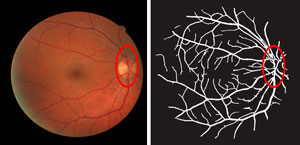
A retinal image (left) and the blood vessel network traced by the ‘absorbing random walk’ image processing algorithm.
© 2017 IEEE. Reprinted, with permission, from Ref 1
More accurate and efficient mapping of retinal blood vessels using a path-following image processing scheme, developed by an A*STAR-led research team, could help improve retinal scanning and medical diagnosis.
The blood vessels found on the retina at the back of the eye are an important diagnostic indicator for many clinical disorders including diabetes, high blood pressure, arterial hardening, and occlusion of retinal arteries. However, tracing retinal blood vessels is a time-consuming process requiring training and skill, which would be better performed by a reliable automated process that can efficiently map the vessel network.
“We have spent years analyzing retinal blood vessels, where a challenge is always to single out each vessel from the rest or to separate artery from vein vessels,” says Cheng Li from the A*STAR Bioinformatics Institute. “We have developed an algorithm that can trace a network from a few marked or ‘labeled’ nodes, and it works especially well for large-scale networks of, say, millions of nodes even with very few known labels.”
In their theoretical study, Li and his team explored the use of a well-established algorithm in image processing, called the Markov chain, to better follow the complex branching networks of blood vessels in the retina.
A Markov chain is a statistical representation of a sequence, in this case of connected nodes, where an element in the sequence is independent of everything that came before it. For a blood vessel, this means that its direction of branching from a given point could be entirely random and not dependent on the path of the vessel that came before it. Li’s team took this further to adopt an absorbing Markov chain, which ‘locks in’ the traced path up to the current node, and then applies a random walk algorithm to probe an image for the next blood vessel direction.
In this way, their image processing algorithm can start from a labeled node, such as a major branch, and trace the blood vessels to form a connected network in a way similar to how a physician would tackle the problem.
In application to real retinal images, the algorithm outperformed other state-of-the-art approaches, and matched the accuracy of human tracing.
“We developed this algorithm out of our very practical biomedical imaging experience in blood vessel tracing over a number of years,” says Li. “Our approach is simple, easy to implement, and has many important applications including image classification, and network and link analysis.”
The A*STAR-affiliated researchers contributing to this research are from the Bioinformatics Institute.



