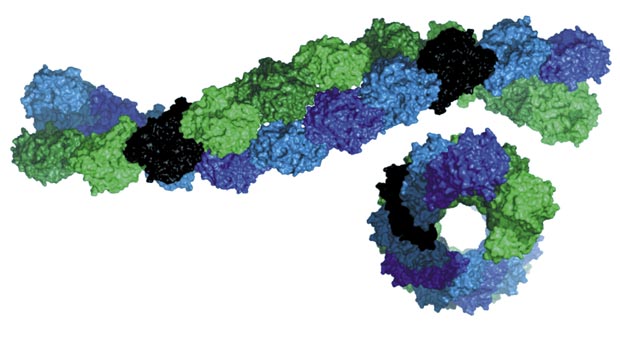
Two strands of the bacterial protein (shown in green and blue) combine to form tubular filaments.
Modified, with permission, from Ref. 1 © 2016 S. Jiang et al.
In both higher organisms and bacteria, DNA must be segregated when cells divide, ensuring that the requisite share of duplicated DNA goes into each new cell. While previous studies indicated that bacteria and higher organisms use quite different systems to perform this task, A*STAR researchers have now found a bacterium that uses filaments with key similarities to those in multicellular organisms, including humans.
Robert Robinson from the A*STAR Institute of Molecular and Cell Biology has a long-standing interest in what he calls the “biological machines” that move DNA around when cells divide. He and his co-workers had gleaned from gene sequencing analysis that there was something distinctive about the DNA-moving machinery in the bacterium Bacillus thuringiensis.
Along with an international team of colleagues, the A*STAR researchers used electron microscopy and biochemical analysis to investigate the way small circular strands of DNA called plasmids moved in this bacterium. They identified a novel form of bacterial filament that combines to form tubules with some similarities to the microtubules observed in higher organisms. Bacterial systems previously studied also use protein filaments to move DNA, but they do not share such obvious similarities to those of higher organisms. The new-found similarity in Bacillus thuringiensis is of great interest from an evolutionary perspective as it suggests that evolution has furnished at least some bacteria and multicellular organisms with different machineries but similar methods to manipulate DNA.

The A*STAR Institute of Molecular and Cell Biology team: (from left to right) Robert Robinson, David Popp and Shimin Jiang.
© 2016 A*STAR Institute of Molecular and Cell Biology
There are practical possibilities for applying this knowledge, beyond its relevance to understanding bacteria. The finding specifically sheds light on the movement of plasmid DNA and genetic engineers use plasmids as vehicles to carry foreign genes into bacteria to modify them to perform useful new tasks. “Understanding plasmid segregation can allow us to interfere with it,” says Robinson. He explains that the research is revealing “new tools” that might be used to manipulate bacteria as well as being applied to synthetic biology — the attempt to construct new living systems from simple parts.
“We are keen to team up with scientists in Singapore who are working to build nanomachines and nanowires,” Robinson adds. The concept of making molecular components and machines for use in manufacturing, medicine or computing is attracting the attention of many research teams worldwide. Thus the revelation found in Bacillus thuringiensis may one day be used to manipulate DNA and other molecules within synthetic structures that extend the abilities of biological machinery into new applications.
The A*STAR-affiliated researchers contributing to this research are from the Institute of Molecular and Cell Biology