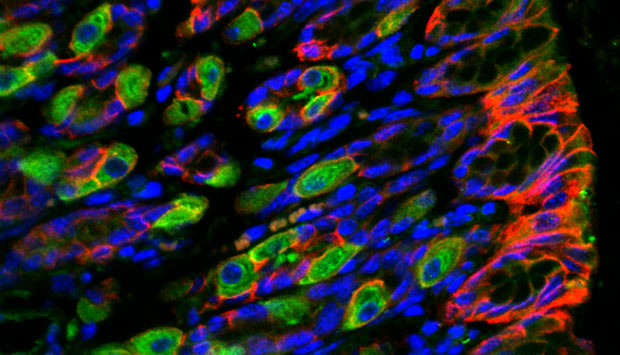
Immunofluorescence image of human stomach section stained for a gene involved in regulating cellular adhesion ARHGAP26 (green), epithelial cadherin (red) and cell nuclei (blue).
© 2015 A*STAR Institute of Molecular and Cell Biology and A*STAR Genome Institute of Singapore
Researchers at A*STAR have identified a genomic re-arrangement linked to the development and spread of gastric cancer.
The DNA in our cells is subject to constant wear and tear from copying mistakes during replication to breaks which sever a chromosome. Extensive repair mechanisms helps cells cope with this damage, but the process is not perfect — when fragments of two broken chromosomes join to form a hybrid chromosome, the genes adjacent to the break sites may fuse. These ‘fusion genes’ are associated with a higher cancer risk.
“I’m amazed what a mutational load every cell in our body has to face every day, and the repair systems and redundancy that let us tolerate so much damage,” says Axel Hillmer of the A*STAR Genome Institute of Singapore, who led the study. “These mechanisms, however, can also result in mutations which cause cancer.”
To identify fusion genes related to gastric cancer, Hillmer’s team compared the genomes of cells from tumors to genomes of normal cells from the same patients. Five of these ‘fusions’ were more common than expected in the gastric cancer samples, suggesting they increase the likelihood of tumor formation.
One of the fusions in the gastric cancer sample combined two genes involved in regulating cellular adhesion, CLDN18 and ARHGAP26. The team used a variety of approaches to investigate the changes wrought by the fusion gene and discovered that it reduces cellular adhesion and aggregation. This weakens the connection between cells in the gastric epithelium, allowing stomach acids to leak out, which leads to chronic gastric inflammation and increases the risk of gastric cancer. Once the cells reach a cancerous state, the fusion gene also enhances their invasiveness and their propensity to metastasize. The fusion therefore plays an important role in both the early and late stages of cancer progression.
“This isn’t a classic oncogene, but it has other features that make it important for cancer. That makes it special — it’s uncommon to find a cancer gene that isn’t an oncogene or a tumor suppressor,” says Walter Hunziker, co-senior author of the study.

Members of Axel Hillmer’s team: Top from left to right: Huay Mei Poh, Rahul Nahar, Yao Fei, Yin Yeng Lee, Royston Kwok, Asif Javed, Kenneth Koh, Say Li Kong, Xingliang Liu. Bottom from left to right: Yee Yen Sia, Joyce Tai, Elaine Chew, Hamizah Abdul Ghani, Jacelyn Phua, Thushangi Pathiraya, Axel Hillmer.
© 2015 A*STAR Genome Institute of Singapore
Unlike oncogenes, which make a cell cancerous, the fusion gene contributes to conditions which are likely to lead to cancer. Though the impact of oncogenes makes them easier to detect and characterize than more subtle players such as these fusion genes — which Hillmer suspects may be more common than realized — both hold promise for cancer diagnosis.
As the CLDN18–ARHGAP26 fusion was not detected in normal gastric tissue, the researchers are hopeful about its potential as an early biomarker for gastric cancer, following further validation.
The A*STAR-affiliated researchers contributing to this research are from the Genome Institute of Singapore and the Institute of Molecular and Cell Biology.